A brief comparison of test equating methods
Magnus Johansson, PhD
Karolinska Institutet, Department of Clinical Neurosciencepgmj@pm.me
2026-02-04
Source:vignettes/articles/comparison.Rmd
comparison.Rmd
library(dplyr)
library(tidyr)
library(ggplot2)
library(leunbachR)
library(equate)
library(SNSequate)
library(knitr)
library(mirt)
library(kequate)
set.seed(1234) # for reproducibility of bootstrap results
select <- dplyr::select
d3a_sum <- read.delim("data/data3a_item.csv", sep = ",") %>%
select(a01:a10,b01:b10) %>%
mutate(a_sum = rowSums(across(c(a01:a10))),
b_sum = rowSums(across(c(b01:b10)))) %>%
select(a_sum,b_sum)
d3a <- read.delim("data/data3a_item.csv", sep = ",") %>%
select(a01:a10,b01:b10)
head(d3a)## a01 a02 a03 a04 a05 a06 a07 a08 a09 a10 b01 b02 b03 b04 b05 b06 b07 b08 b09
## 1 0 0 0 1 0 0 0 0 0 0 1 1 0 0 0 0 1 0 0
## 2 1 1 0 0 1 0 0 1 0 0 1 1 1 1 0 0 0 0 0
## 3 1 0 1 0 1 1 0 0 0 0 1 1 1 1 1 1 1 1 1
## 4 1 1 1 1 1 0 1 0 0 0 1 1 1 1 1 1 0 0 0
## 5 1 1 1 1 1 1 1 1 0 0 1 1 1 0 1 0 1 1 0
## 6 1 1 1 0 1 1 1 0 0 0 1 1 1 1 1 0 0 1 1
## b10
## 1 0
## 2 0
## 3 0
## 4 0
## 5 0
## 6 1While the Leunbach method allows for test equating based on observed sum scores under the assumption that the underlying data fits a Rasch model adequately, most other methods require item-level response data.
We will first focus on comparing IRT true score equating to Leunbach, but also include some other methods.
Direct equating
mod1 <- mirt(d3a[,1:10], 1, 'Rasch', verbose = FALSE)
mod2 <- mirt(d3a[,11:20], 1, 'Rasch', verbose = FALSE)
coef_ls1 <- list(a = rep(1,10),
b = coef(mod1, simplify=TRUE, IRTpars = TRUE)$item[,'b'],
c = rep(0,10))
coef_ls2 <- list(a = rep(1,10),
b = coef(mod2, simplify=TRUE, IRTpars = TRUE)$item[,'b'],
c = rep(0,10))
req1 <- irt.eq(n_items = 10, coef_ls1, coef_ls2, method="TS", A=1, B=0) # true score
rx <- freqtab(rowSums(d3a[,1:10]), scales = 0:10)
ry <- freqtab(rowSums(d3a[,11:20]), scales = 0:10)
req2 <- equate(rx, ry, type = "i") # identity
req3 <- equate(rx, ry, type = "l") # linear
req4 <- equate(rx, ry, type = "e", smooth = "loglin", degrees = 3) # equipercentile with loglinear smoothing
# Leunbach model
lfit <- leunbach_ipf(d3a_sum)
lboot <- leunbach_bootstrap(lfit, n_cores = 4, nsim = 100)
leq <- get_equating_table(lboot)
# summary table
max_score <- 10
eq_table <- data.frame(identity = pmin(pmax(req2$conc$yx, 0), max_score),
linear = pmin(pmax(req3$conc$yx, 0), max_score),
equiperc_loglin3 = pmin(pmax(req4$conc$yx, 0), max_score),
IRT_truescore = req1$tau_y,
leunbach_expected = leq$expected,
IRT_thetaequivalent = req1$theta_equivalent,
leunbach_theta = leq$log_theta
) %>%
round(2)
kable(eq_table)| identity | linear | equiperc_loglin3 | IRT_truescore | leunbach_expected | IRT_thetaequivalent | leunbach_theta |
|---|---|---|---|---|---|---|
| 0 | 0.36 | 0.00 | 0.00 | 0.00 | NA | -5.00 |
| 1 | 1.30 | 0.96 | 0.91 | 0.79 | -2.18 | -1.98 |
| 2 | 2.23 | 2.10 | 1.93 | 1.90 | -1.49 | -1.33 |
| 3 | 3.16 | 3.19 | 2.99 | 3.07 | -0.94 | -0.86 |
| 4 | 4.10 | 4.20 | 4.04 | 4.16 | -0.45 | -0.45 |
| 5 | 5.03 | 5.14 | 5.08 | 5.11 | 0.02 | -0.07 |
| 6 | 5.97 | 6.03 | 6.10 | 5.95 | 0.49 | 0.29 |
| 7 | 6.90 | 6.88 | 7.09 | 6.71 | 0.98 | 0.66 |
| 8 | 7.84 | 7.64 | 8.08 | 7.53 | 1.51 | 1.13 |
| 9 | 8.77 | 8.42 | 9.06 | 8.80 | 2.19 | 2.01 |
| 10 | 9.71 | 9.38 | 10.00 | 10.00 | NA | 5.00 |
eq_table %>%
select(!c(leunbach_theta,IRT_thetaequivalent)) %>%
pivot_longer(!identity) %>%
ggplot(aes(x = identity, y = value, color = name, shape = name, linetype = name)) +
geom_point(size = 2.5) +
geom_line() +
scale_x_continuous(breaks = c(0:10), minor_breaks = NULL) +
scale_y_continuous(breaks = c(0:10)) +
scale_color_viridis_d() +
theme_bw() +
labs(title = "Test equating on sum score scale",
color = "Equating model", shape = "Equating model", linetype = "Equating model",
y = "Predicted Y value", x = "X value")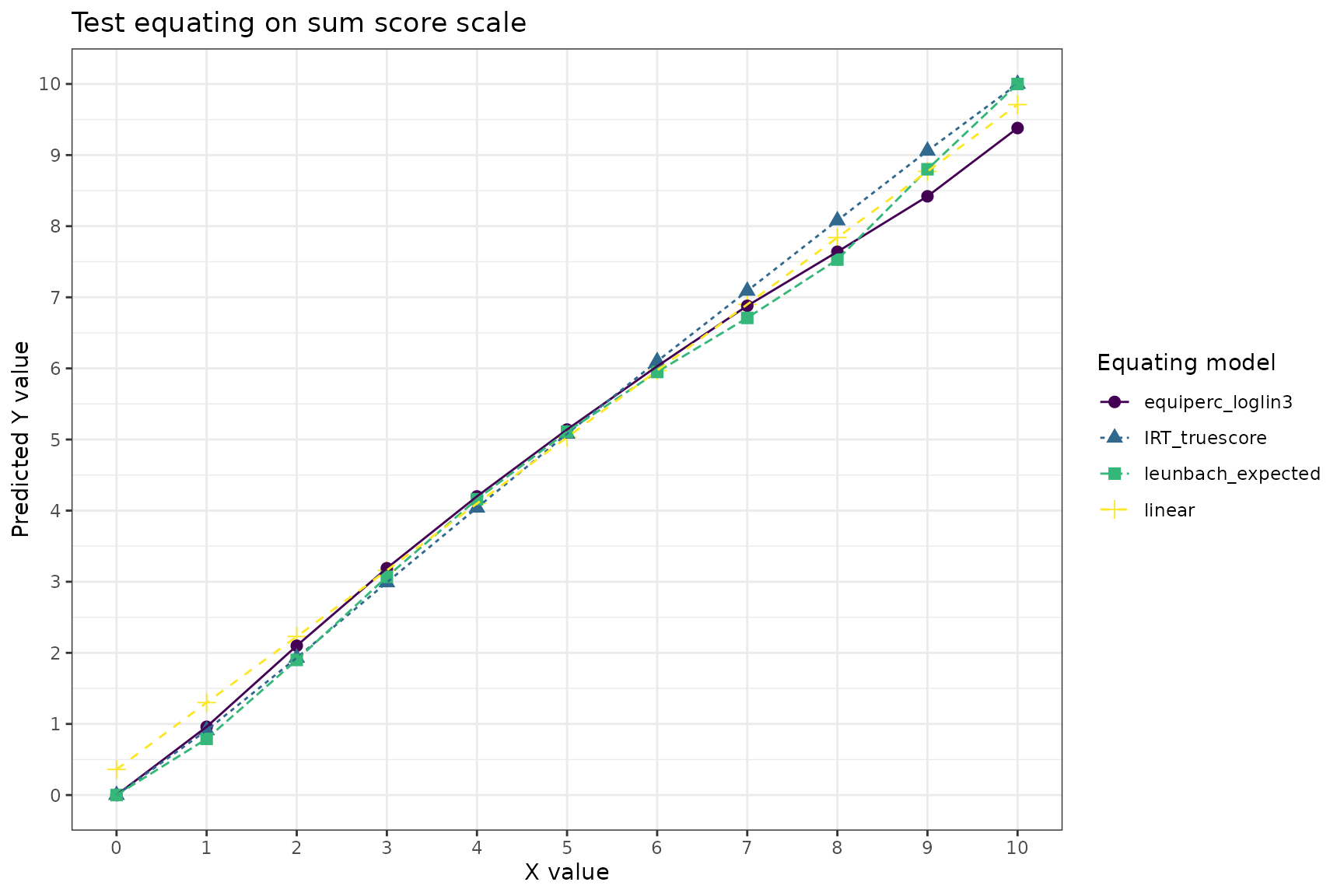
eq_table %>%
select(c(identity,leunbach_theta,IRT_thetaequivalent)) %>%
pivot_longer(!identity) %>%
filter(identity %in% c(1:9)) %>%
ggplot(aes(x = identity, y = value, color = name, shape = name)) +
geom_point(size = 3) +
geom_line() +
scale_x_continuous(breaks = c(0:10), minor_breaks = NULL) +
scale_y_continuous(breaks = c(-2:2)) +
scale_color_viridis_d() +
theme_bw() +
labs(title = "Test equating on theta scale",
color = "Equating model", shape = "Equating model",
y = "Predicted Y value", x = "X value", caption = "Note: Min and max scores are excluded, since these need to be imputed for both methods.") +
theme(plot.caption = element_text(face = "italic", hjust = 0))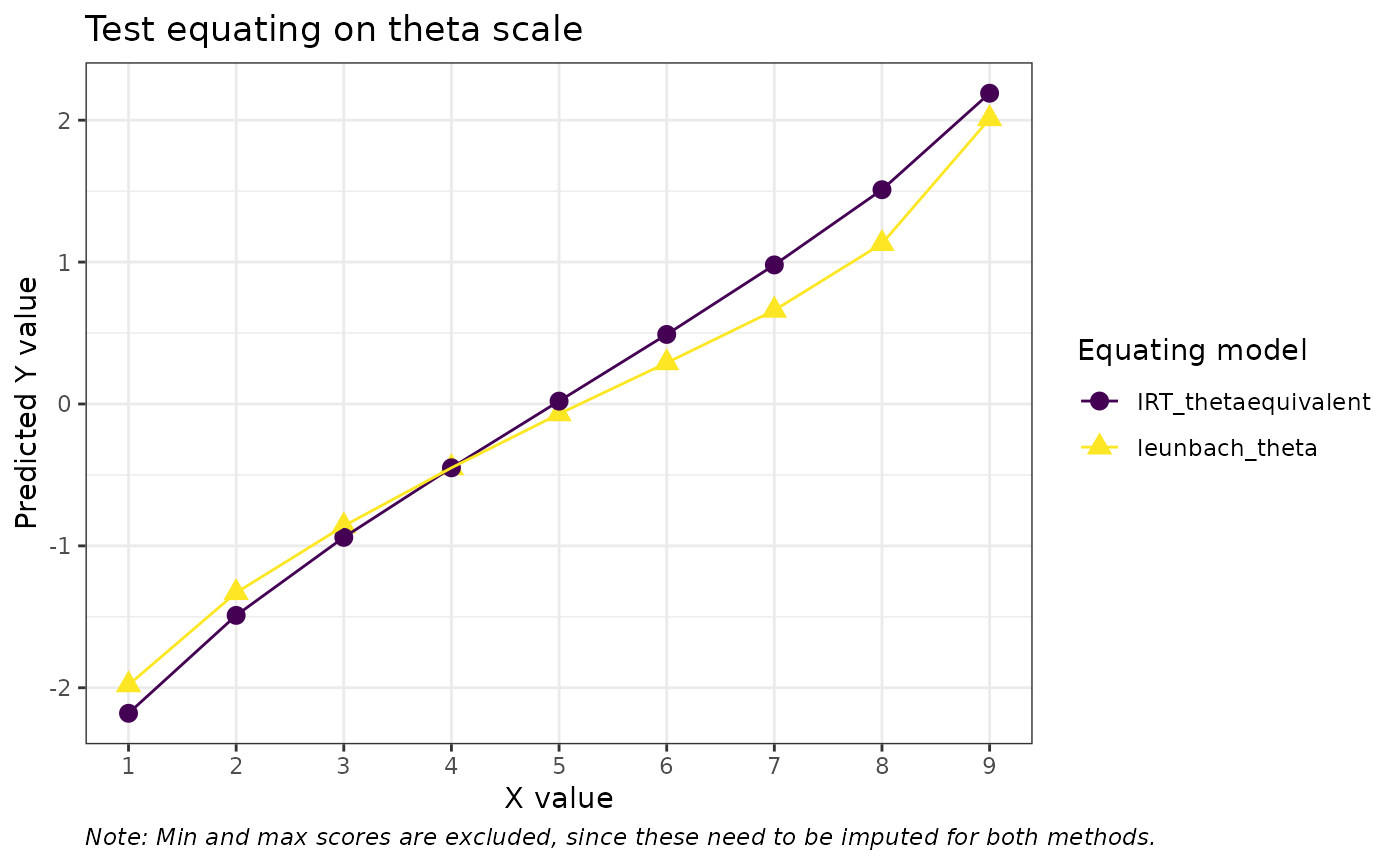
Kernel equating
We’ll apply the equivalent groups design, following the
kequate package vignette (Andersson
et al. 2022). First, we fit two separate generalized linear
models (GLM) using the poisson distribution for the counts of each
score. The kequate vignette suggests using AIC to evaluate
model fit (lower values are better) and finding the optimal number of
moments to include in the model specification. I have opted for using
B-splines instead for slightly improved model fit, and adjusting the
degrees of freedom based on AIC. The glm() output objects
are then used by the kequate() function.
Notably, like Leunbach, kernel equating also allows for equating using only sum scores.
glm_a <- glm(count ~ splines::bs(total, df = 3),
family = "poisson", data = rx, x = TRUE)
glm_b <- glm(count ~ splines::bs(total, df = 3),
family = "poisson", data = ry, x = TRUE)Bias correction following Kosmidis et al. (2020).
library(brglm2)
glm_a2 <- update(glm_a, method = "brglmFit")
glm_b2 <- update(glm_b, method = "brglmFit")
eg_eq <- kequate(
design = "EG",
r = glm_a2, s = glm_b2,
x = c(0:10), y = c(0:10)
)
data.frame(
predicted = c(eg_eq@equating$eqYx, leq$expected),
Model = c(rep("Kernel equating",11),rep("Leunbach",11)),
sumscore = c(0:10,0:10)
) %>%
ggplot(aes(x = sumscore, y = predicted, color = Model, shape = Model)) +
geom_point(size = 3) +
geom_line() +
scale_x_continuous(breaks = c(0:10), minor_breaks = NULL) +
#scale_y_continuous(breaks = c(-2:2)) +
scale_color_viridis_d() +
theme_bw() +
labs(title = "Test equating expected scores",
color = "Equating model", shape = "Equating model",
y = "Predicted Y value", x = "X value") +
theme(plot.caption = element_text(face = "italic", hjust = 0))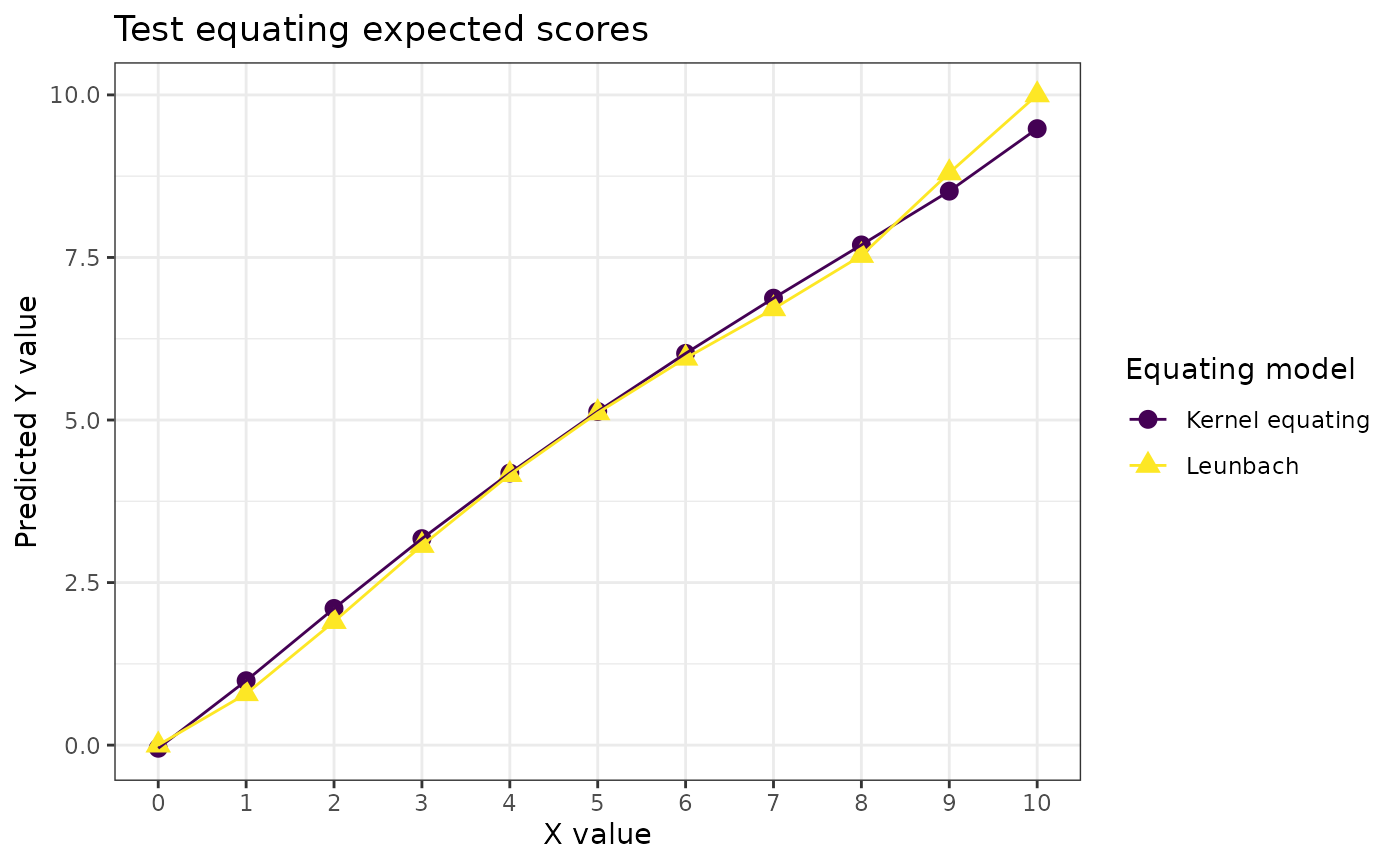
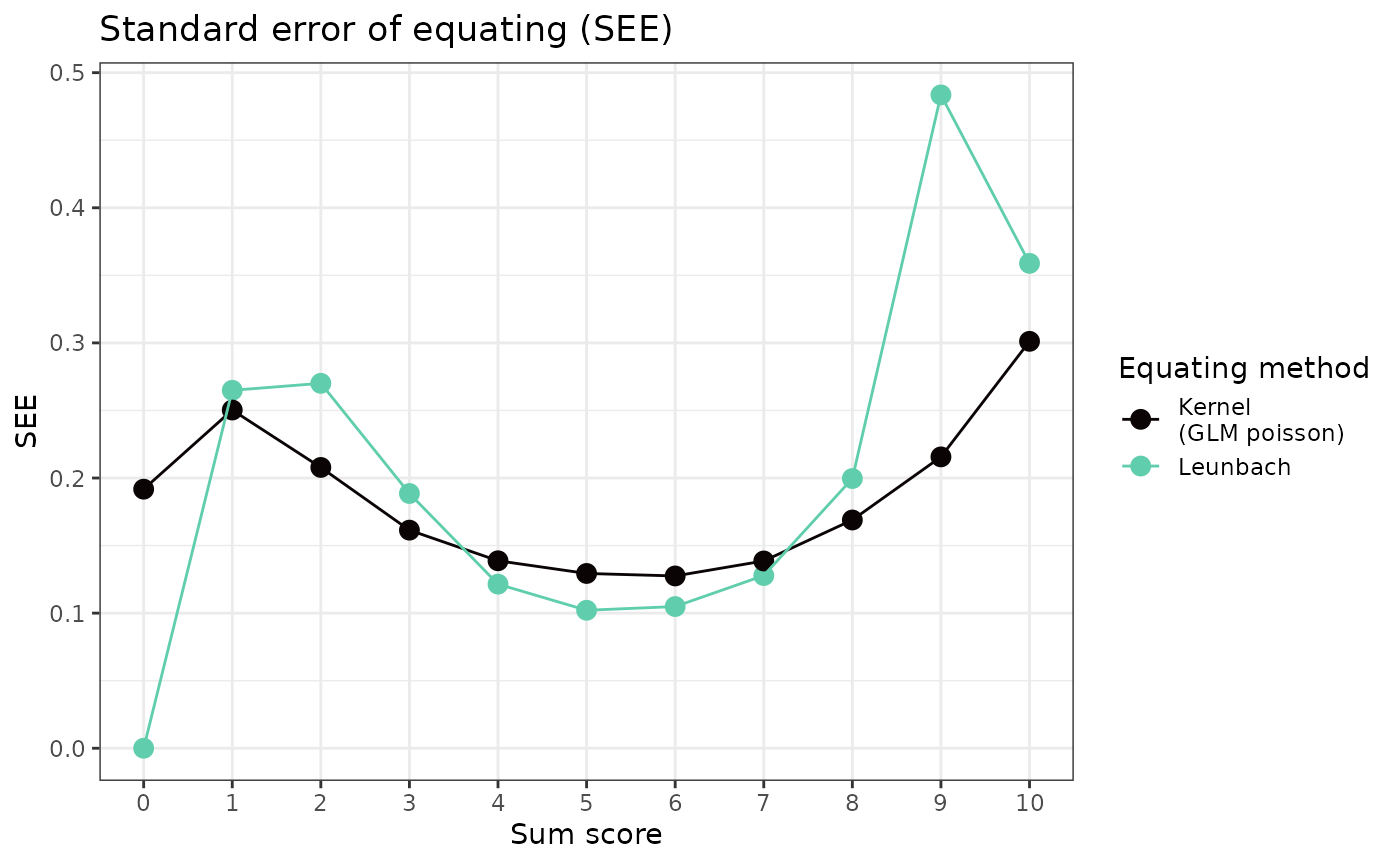
Standard error of equating
In order to make reasonable comparisons between kernel equating and Leunbach, we need to use the expected values for estimating standard errors of equating (SEE) in the Leunbach bootstrap, rather than the rounded values (default). It should be noted that the methods of estimating SEE are different for these two equating models.
lboot2 <- leunbach_bootstrap(lfit, n_cores = 4, nsim = 100, see_type = "expected")
data.frame(score = c(0:10,0:10),
see = c(lboot2[["see_1to2"]],eg_eq@equating$SEEYx),
model = c(rep("Leunbach",11),rep("Kernel\n(GLM poisson)",11))
) %>% ggplot(aes(x=score,y=see, color = model)) +
geom_point(size = 3) +
geom_line() +
theme_bw() +
scale_x_continuous(breaks = c(0:10), minor_breaks = NULL) +
scale_color_viridis_d(option = "G", end = 0.8) +
labs(title = "Standard error of equating (SEE)",
x = "Sum score", y = "SEE", color = "Equating method")